A newly published study in the journal Gastroenterology by Peter Rimmer, Fan Zhang, and colleagues digs into the observation that there is a shift in the microbiome in newly diagnosed patients with irritable bowel disease. The two most common IBD types are Crohn’s disease and ulcerative colitis, with the global prevalence exceeing 0.3%. That’s nearly 2.5 million people!
The underlying causes of IBD are not clear, but there seems to be an interplay between several contributors: barrier dysfunction, genetic predisposition, environmental factors, and the gut microbiome. Alterations in the gut bacteria in IBD patients have been widely reported. This leads to a breakdown of homeostasis, leading to an increase in pathogenic bacteria activity and inappropriate immune responses. But the question remains: is the microbiome alteration a cause or a symptom of IBD? This paper is unique in that it was a meta-analysis that examined 36 studies, but redid the bioinformatic analysis, where possible, to create consistency across the data sets. Visualizing bioinformatic data is a complex task. Bioinformatic analysis is an important step because the choices made in the analysis of the raw bioinformatic data can alter interpretations. Adding a layer of a new, consistent analysis made interpretations easier.
I think the key finding was the demonstration of a depletion of anaerobes and an enrichment in oxygen-tolerant bacteria in newly diagnosed IBD patients. (See the reproduction of Figure 3A below)The study also found a decrease in diversity in IBD patients, in agreement with other research. Another interesting finding of the analyzed data was that the microbiome changes depending on the person’s location. Some samples from North America were significantly different from those from Asia or Africa, but the disease state was far more important.
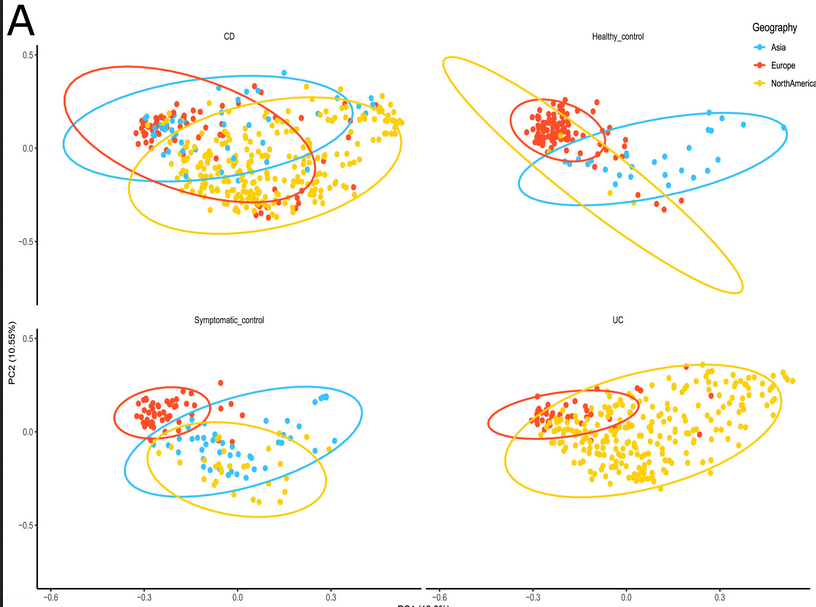
This deeper understanding of the shifts in the microbiome caused by IBD may lead to interventions that help move IBD patients back to a healthier microbiome.